Curvas PES de dissociação com Qunova HiVQE
As Funções Qiskit são um recurso experimental disponível apenas para usuários dos Planos Premium, Flex e On-Prem (via API da Plataforma IBM Quantum) do IBM Quantum®. Elas estão em status de versão prévia e sujeitas a alterações.
Estimativa de uso (NOTA: Esta é apenas uma estimativa. Seu tempo de execução pode variar.)
- Li2S: Cinco minutos de tempo de QPU em um processador Heron r2
- FeP-NO: Cinco minutos de tempo de QPU em um processador Heron r2
Contexto
Calcular com precisão as energias de reações químicas é crucial para avanços científicos em ciência dos materiais, engenharia química, descoberta de medicamentos e outros campos. Entre vários sistemas químicos, o sistema Li-S tem despertado interesse significativo para compreender e desenvolver novas composições de baterias. Este tutorial fornece experiência prática no cálculo da superfície de energia potencial (PES) de dissociação da ligação Li-S de um sistema removendo um átomo de lítio usando cálculos HiVQE. Os resultados podem ser comparados com cálculos de referência (CASCI), bem como métodos clássicos como Hartree-Fock (HF) para um problema de 20 qubits.
Requisitos
Instale as seguintes dependências para executar o código neste tutorial.
!pip install --upgrade pip
!pip install -U qiskit-ibm-catalog "qiskit_ibm_runtime<0.42.0" pyscf numpy matplotlib typing_extensions
Configuração
Para executar este tutorial, importe a função qunova/hivqe-chemistry via QiskitFunctionCatalog. Você precisa de uma conta com Plano Premium, Plano Flex ou Plano On-Prem (API da Plataforma IBM Quantum) do IBM Quantum com uma licença da Qunova para executar esta função.
from qiskit_ibm_catalog import QiskitFunctionsCatalog
from pyscf import gto, scf, mcscf
import matplotlib.pyplot as plt
import pprint
catalog = QiskitFunctionsCatalog(
channel="ibm_quantum_platform",
instance="INSTANCE_CRN",
token="YOUR_API_KEY", # Use the 44-character API_KEY you created and saved from the IBM Quantum Platform Home dashboard
)
hivqe = catalog.load("qunova/hivqe-chemistry")
Parte 1: Li2S (20Q)
Passo 1: Mapear entradas clássicas para um problema quântico
Defina as geometrias de em formato de dicionário para diferentes distâncias de ligação Li-S para calcular a curva PES. Estas geometrias são otimizadas usando cálculos B3LYP/631g.
str_geometries = {
"1.51": "S -1.239044 0.671232 -0.030374; Li -1.506327 0.432403 -1.498949; Li -0.899996 0.973348 1.826768",
"1.91": "S -1.215858 0.692272 0.099232; Li -1.553305 0.390283 -1.758043; Li -0.876205 0.994426 1.956257",
"2.40": "S -1.741432 0.680397 0.346702; Li -0.529307 0.488006 -1.729343; Li -1.284307 0.989409 2.177209",
"3.10": "S -2.347450 0.657089 0.566194; Li -0.199353 0.527517 -1.665148; Li -1.008243 0.973206 1.893522",
"3.80": "S -2.707255 0.674298 0.909161; Li 0.079218 0.552012 -1.671656; Li -0.927010 0.931502 1.557063",
"4.50": "S -2.913363 0.709175 1.276987; Li 0.368656 0.559989 -1.798088; Li -1.010340 0.888647 1.315670",
}
str_geometries
{'1.51': 'S -1.239044 0.671232 -0.030374; Li -1.506327 0.432403 -1.498949; Li -0.899996 0.973348 1.826768',
'1.91': 'S -1.215858 0.692272 0.099232; Li -1.553305 0.390283 -1.758043; Li -0.876205 0.994426 1.956257',
'2.40': 'S -1.741432 0.680397 0.346702; Li -0.529307 0.488006 -1.729343; Li -1.284307 0.989409 2.177209',
'3.10': 'S -2.347450 0.657089 0.566194; Li -0.199353 0.527517 -1.665148; Li -1.008243 0.973206 1.893522',
'3.80': 'S -2.707255 0.674298 0.909161; Li 0.079218 0.552012 -1.671656; Li -0.927010 0.931502 1.557063',
'4.50': 'S -2.913363 0.709175 1.276987; Li 0.368656 0.559989 -1.798088; Li -1.010340 0.888647 1.315670'}
Os cálculos HiVQE serão realizados com as opções definidas abaixo. Usando a base sto3g para , há 19 orbitais espaciais com 22 elétrons. Para executar o caso (10o,10e) com cálculo HiVQE, você pode definir 10 orbitais ativos e seis orbitais congelados. Em cada iteração, 100 shots serão usados para amostrar a configuração eletrônica gerada pelo circuito quântico ExcitationPreserving (epa) com entrelaçamento circular e duas repetições (reps). O número máximo de iterações é definido como 30 para garantir o término da iteração com convergência de energia.
molecule_options = {
"basis": "sto3g",
"active_orbitals": list(range(5, 15)),
"frozen_orbitals": list(range(5)),
}
hivqe_options = {
"shots": 100,
"max_iter": 30,
"ansatz": "epa",
"ansatz_entanglement": "circular",
"ansatz_reps": 2,
}
Passos 2 e 3: Otimizar o problema para execução em hardware quântico e executar usando a função HiVQE Chemistry
Configure o loop for para executar cálculos HiVQE com geometrias com as opções definidas abaixo. Os trabalhos são submetidos no loop for. Neste tutorial, você submeterá seis geometrias e recuperará os resultados quando todos estiverem concluídos. Na execução da função principal, você precisa definir max_states e max_expansion_states para controlar o tamanho máximo da matriz do subespaço e para controlar quantos estados podem ser gerados usando métodos de expansão CI clássicos por iteração. Os IDs de trabalho da função serão armazenados no dicionário com cada rótulo de geometria para rastrear e processar ainda mais a saída.
info_jobid = {}
for dis, geom in str_geometries.items():
hivqe_run = hivqe.run(
geometry=geom,
backend_name="",
max_states=40000,
max_expansion_states=100,
molecule_options=molecule_options,
hivqe_options=hivqe_options,
)
status = hivqe_run.status()
info_jobid[dis] = hivqe_run.job_id
print(info_jobid)
{'1.51': 'de3b8818-c9db-4fa3-a3c2-d51551c2dfaf', '1.91': '55d9467a-fc85-49a8-9bc6-8f6990e421e5', '2.40': '415112b3-69ff-4d53-8b10-cb4e3be68c9e', '3.10': 'ef67b600-3887-4225-b872-e354dfdf8454', '3.80': 'b16d3502-a9e4-4560-9775-852e9d07e70f', '4.50': '0c0bffc7-af77-4a56-a656-2a2610c991d6'}
Vamos verificar se todos os trabalhos ainda estão em execução ou foram concluídos.
completed_jobs_num = 0
running_jobs_num = 0
completed_jobs = {}
for i, info in enumerate(info_jobid.items()):
dis, job_id = info
submitted_job = catalog.get_job_by_id(job_id)
stat = submitted_job.status()
print(dis, submitted_job.job_id, stat)
if stat == "DONE":
completed_jobs_num += 1
completed_jobs[dis] = submitted_job
if (stat == "RUNNING") or (stat == "QUEUED"):
running_jobs_num += 1
print(
f"Completed {completed_jobs_num} job, Running or Queued {running_jobs_num} job"
)
1.51 de3b8818-c9db-4fa3-a3c2-d51551c2dfaf DONE
1.91 55d9467a-fc85-49a8-9bc6-8f6990e421e5 DONE
2.40 415112b3-69ff-4d53-8b10-cb4e3be68c9e DONE
3.10 ef67b600-3887-4225-b872-e354dfdf8454 DONE
3.80 b16d3502-a9e4-4560-9775-852e9d07e70f DONE
4.50 0c0bffc7-af77-4a56-a656-2a2610c991d6 DONE
Completed 6 job, Running or Queued 0 job
Uma vez que todos os trabalhos estejam concluídos, vamos recuperar todos os resultados dos cálculos.
hivqe_result = {}
if len(info_jobid) == completed_jobs_num:
print("All jobs are completed")
for i, job in enumerate(completed_jobs.items()):
dis, cal = job
print(dis, cal.result()["energy"])
hivqe_result[str(dis)] = cal.result()["energy"]
All jobs are completed
1.51 -407.8944801731773
1.91 -407.9800570932916
2.40 -407.9372992999806
3.10 -407.86278336000134
3.80 -407.83092972296157
4.50 -407.82971011225766
pprint.pprint(hivqe_result)
{'1.51': -407.8944801731773,
'1.91': -407.9800570932916,
'2.40': -407.9372992999806,
'3.10': -407.86278336000134,
'3.80': -407.83092972296157,
'4.50': -407.82971011225766}
O tempo total de execução da QPU usado no trabalho pode ser rastreado fazendo login na Plataforma IBM Quantum e visualizando os trabalhos submetidos com a tag qunova-chemistry-hivqe.
Passo 4: Pós-processar e comparar com métodos clássicos
O cálculo de referência clássico (CASCI) pode ser conduzido para (10o,10e) para validar os resultados do HiVQE.
str_geometries = {
"1.31": "S -1.250686 0.660708 -0.095168; Li -1.482812 0.453464 -1.369406; Li -0.911870 0.962810 1.762020",
"1.41": "S -1.244856 0.665971 -0.062773; Li -1.494574 0.442933 -1.434177; Li -0.905937 0.968078 1.794395",
"1.51": "S -1.239044 0.671232 -0.030374; Li -1.506327 0.432403 -1.498949; Li -0.899996 0.973348 1.826768",
"1.61": "S -1.233245 0.676492 0.002027; Li -1.518073 0.421873 -1.563722; Li -0.894049 0.978617 1.859141",
"1.71": "S -1.227453 0.681752 0.034429; Li -1.529816 0.411343 -1.628496; Li -0.888099 0.983887 1.891513",
"1.81": "S -1.221659 0.687012 0.066831; Li -1.541558 0.400813 -1.693270; Li -0.882150 0.989157 1.923885",
"1.91": "S -1.215858 0.692272 0.099232; Li -1.553305 0.390283 -1.758043; Li -0.876205 0.994426 1.956257",
"2.01": "S -1.209887 0.697544 0.131599; Li -1.565136 0.379748 -1.822800; Li -0.870344 0.999691 1.988646",
"2.11": "S -1.203945 0.702813 0.163973; Li -1.576953 0.369214 -1.887560; Li -0.864469 1.004956 2.021033",
"2.21": "S -1.198023 0.708081 0.196350; Li -1.588760 0.358680 -1.952322; Li -0.858584 1.010221 2.053417",
"2.30": "S -1.365426 0.717714 0.367060; Li -0.689401 0.458925 -1.828368; Li -1.500219 0.981173 2.255876",
"2.31": "S -1.192118 0.713348 0.228731; Li -1.600559 0.348146 -2.017085; Li -0.852690 1.015488 2.085800",
"2.40": "S -1.741432 0.680397 0.346702; Li -0.529307 0.488006 -1.729343; Li -1.284307 0.989409 2.177209",
"2.50": "S -1.885961 0.669986 0.365815; Li -0.461563 0.499084 -1.695846; Li -1.207523 0.988741 2.124599",
"2.60": "S -1.977163 0.665155 0.389784; Li -0.416654 0.504966 -1.683655; Li -1.161229 0.987690 2.088439",
"2.70": "S -2.063642 0.661518 0.418977; Li -0.367600 0.510505 -1.676408; Li -1.123804 0.985788 2.051998",
"2.80": "S -2.141072 0.659218 0.451663; Li -0.323153 0.515056 -1.673046; Li -1.090821 0.983538 2.015951",
"2.90": "S -2.212097 0.657968 0.487535; Li -0.281989 0.518909 -1.672407; Li -1.060960 0.980935 1.979440",
"3.00": "S -2.281477 0.657123 0.525155; Li -0.239607 0.523326 -1.668669; Li -1.033963 0.977363 1.938081",
"3.10": "S -2.347450 0.657089 0.566194; Li -0.199353 0.527517 -1.665148; Li -1.008243 0.973206 1.893522",
"3.20": "S -2.410882 0.657532 0.608912; Li -0.157788 0.532069 -1.659971; Li -0.986376 0.968211 1.845627",
"3.30": "S -2.470306 0.658818 0.654893; Li -0.118007 0.536237 -1.656311; Li -0.966733 0.962757 1.795986",
"3.40": "S -2.525776 0.660762 0.702910; Li -0.078312 0.540189 -1.654076; Li -0.950958 0.956861 1.745734",
"3.50": "S -2.576885 0.663376 0.752788; Li -0.039076 0.543706 -1.654536; Li -0.939085 0.950730 1.696316",
"3.60": "S -2.623930 0.666534 0.803853; Li 0.000274 0.546839 -1.657697; Li -0.931390 0.944439 1.648412",
"3.70": "S -2.667364 0.670217 0.856250; Li 0.039572 0.549616 -1.663265; Li -0.927254 0.937980 1.601583",
"3.80": "S -2.707255 0.674298 0.909161; Li 0.079218 0.552012 -1.671656; Li -0.927010 0.931502 1.557063",
"3.90": "S -2.744005 0.678718 0.962425; Li 0.119268 0.554073 -1.682595; Li -0.930310 0.925021 1.514738",
"4.00": "S -2.777891 0.683415 1.015798; Li 0.159751 0.555810 -1.696024; Li -0.936907 0.918587 1.474794",
"4.10": "S -2.809179 0.688333 1.069057; Li 0.200678 0.557234 -1.711873; Li -0.946546 0.912245 1.437385",
"4.20": "S -2.838194 0.693443 1.122205; Li 0.242066 0.558401 -1.729770; Li -0.958918 0.905968 1.402134",
"4.30": "S -2.864984 0.698619 1.174415; Li 0.283858 0.559186 -1.750539; Li -0.973920 0.900007 1.370693",
"4.40": "S -2.889984 0.703887 1.226140; Li 0.326068 0.559728 -1.773231; Li -0.991131 0.894196 1.341660",
"4.50": "S -2.913363 0.709175 1.276987; Li 0.368656 0.559989 -1.798088; Li -1.010340 0.888647 1.315670",
}
rhf_result = {}
casci_result = {}
cas_list = molecule_options["active_orbitals"]
distance_ref = []
for dis, geom in str_geometries.items():
distance_ref.append(dis)
mole = gto.M(atom=geom, basis=molecule_options["basis"])
mole.verbose = 0
# RHF energy
mf = scf.RHF(mole).run()
mo_occ = mf.mo_occ
num_elecs_as = int(sum([mo_occ[idx] for idx in cas_list]))
rhf_result[str(dis)] = mf.e_tot
# CASCI energy
casci_solver = mcscf.CASCI(mf, len(cas_list), num_elecs_as)
orbs = mcscf.addons.sort_mo(casci_solver, mf.mo_coeff, cas_list, base=0)
casci_solver.kernel(orbs)
casci_result[str(dis)] = casci_solver.e_tot
print(
f"d={dis:4.3} RHF Energy: {mf.e_tot:14.10}, CASCI Energy: {casci_solver.e_tot:14.10}"
)
d=1.3 RHF Energy: -407.7137006, CASCI Energy: -407.7193917
d=1.4 RHF Energy: -407.8183196, CASCI Energy: -407.8245211
d=1.5 RHF Energy: -407.8878013, CASCI Energy: -407.8944802
d=1.6 RHF Energy: -407.9315356, CASCI Energy: -407.9385663
d=1.7 RHF Energy: -407.9569034, CASCI Energy: -407.9641258
d=1.8 RHF Energy: -407.9693681, CASCI Energy: -407.9766313
d=1.9 RHF Energy: -407.9728592, CASCI Energy: -407.9800572
d=2.0 RHF Energy: -407.9701684, CASCI Energy: -407.9772549
d=2.1 RHF Energy: -407.9632701, CASCI Energy: -407.9702381
d=2.2 RHF Energy: -407.9535584, CASCI Energy: -407.9604007
d=2.3 RHF Energy: -407.9420173, CASCI Energy: -407.9487043
d=2.3 RHF Energy: -407.9420156, CASCI Energy: -407.9487024
d=2.4 RHF Energy: -407.9297216, CASCI Energy: -407.9372993
d=2.5 RHF Energy: -407.9172, CASCI Energy: -407.9261859
d=2.6 RHF Energy: -407.9061139, CASCI Energy: -407.915961
d=2.7 RHF Energy: -407.8937118, CASCI Energy: -407.904259
d=2.8 RHF Energy: -407.8816389, CASCI Energy: -407.8928292
d=2.9 RHF Energy: -407.8700448, CASCI Energy: -407.8819574
d=3.0 RHF Energy: -407.859054, CASCI Energy: -407.8719092
d=3.1 RHF Energy: -407.8487619, CASCI Energy: -407.8628304
d=3.2 RHF Energy: -407.8392304, CASCI Energy: -407.8548482
d=3.3 RHF Energy: -407.8304842, CASCI Energy: -407.8480217
d=3.4 RHF Energy: -407.8225124, CASCI Energy: -407.8423743
d=3.5 RHF Energy: -407.8152758, CASCI Energy: -407.8378892
d=3.6 RHF Energy: -407.8087161, CASCI Energy: -407.8345331
d=3.7 RHF Energy: -407.802764, CASCI Energy: -407.8322563
d=3.8 RHF Energy: -407.7973458, CASCI Energy: -407.83093
d=3.9 RHF Energy: -407.7923883, CASCI Energy: -407.8303555
d=4.0 RHF Energy: -407.7878216, CASCI Energy: -407.83025
d=4.1 RHF Energy: -407.783582, CASCI Energy: -407.8303243
d=4.2 RHF Energy: -407.7796124, CASCI Energy: -407.8303791
d=4.3 RHF Energy: -407.7758633, CASCI Energy: -407.8302885
d=4.4 RHF Energy: -407.7722923, CASCI Energy: -407.8300614
d=4.5 RHF Energy: -407.7688641, CASCI Energy: -407.829711
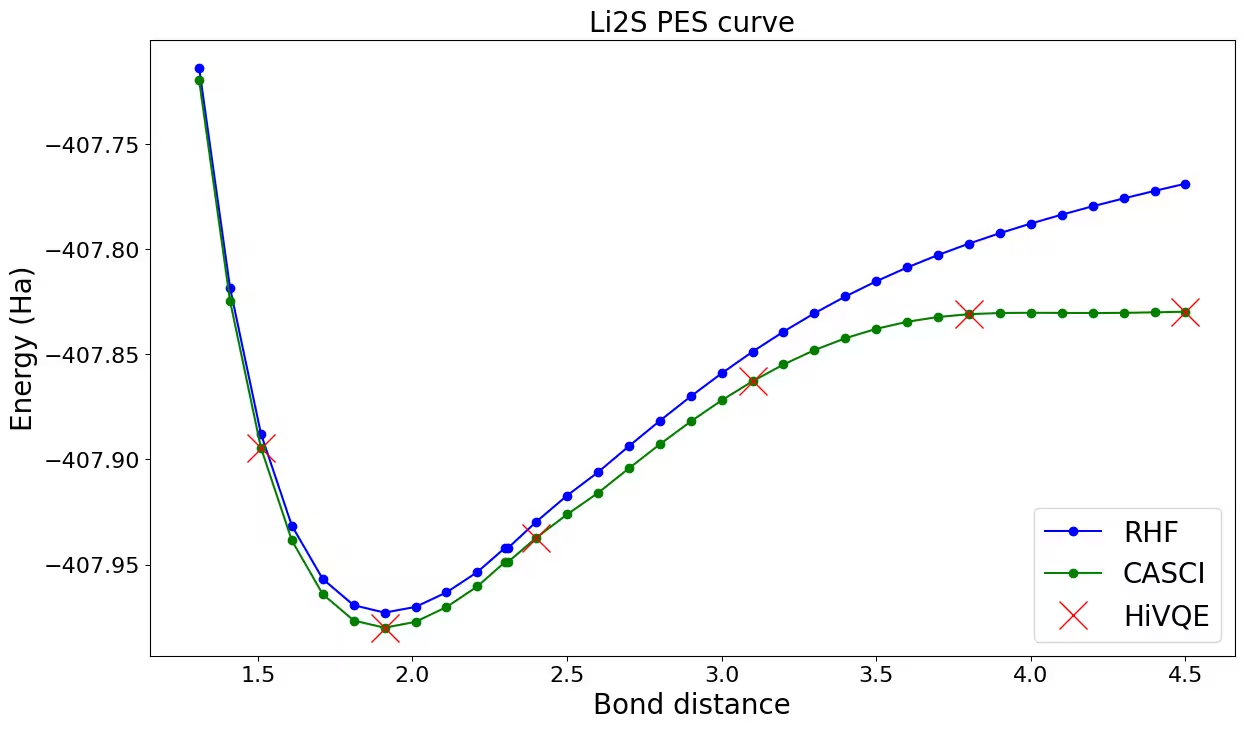
Plotando a curva de dissociação para Li_2S
Vamos plotar e comparar os resultados do HiVQE com HF e CASCI. Você pode observar que todos os cálculos HiVQE estão bem alinhados com o resultado de referência clássico (CASCI).
fig, ax = plt.subplots(1, 1)
hf_energy = [v for key, v in rhf_result.items()]
casci_energy = [v for key, v in casci_result.items()]
hivqe_energy = [v for key, v in hivqe_result.items()]
distance_ref = [float(key) for key, v in rhf_result.items()]
distance = [float(key) for key, v in hivqe_result.items()]
ax.plot(distance_ref, hf_energy, "-o", label="RHF", c="blue")
ax.plot(distance_ref, casci_energy, "-o", label="CASCI", c="green")
ax.plot(distance, hivqe_energy, "x", label="HiVQE", c="red", markersize=20)
ax.legend(fontsize=20)
ax.tick_params("both", labelsize=16)
ax.set_xlabel("Bond distance (angstrom)", size=20)
ax.set_ylabel("Energy (Ha)", size=20)
ax.set_title("Li2S PES curve", size=20)
fig.set_size_inches(14, 8)
Parte 2: FeP-NO (44Q)
Observe que, para executar a função no sistema FeP-NO com as configurações mostradas neste exemplo, você precisa de uma licença que permita usar a função com pelo menos 44 qubits. Envie um e-mail para qiskit.support@qunovacomputing.com para consultar sobre como obter uma licença.
Passo 1: Mapear entradas clássicas para um problema quântico
Defina as opções para os cálculos HiVQE
molecule_options = {
"basis": "631g*",
"active_orbitals": list(range(90, 112, 1)),
"frozen_orbitals": list(range(0, 90, 1)),
"charge": -1,
}
hivqe_options = {
"shots": 2000,
"max_iter": 40,
"ansatz": "epa",
"ansatz_entanglement": "linear",
"ansatz_reps": 2,
"amplitude_screening_tolerance": 1e-6,
}
Defina as geometrias FeP-NO em formato de dicionário para diferentes distâncias de ligação Fe-N para calcular a curva PES.
geometry_1_75 = """
Fe 9.910596 31.534095 1.798088
N 10.557481 31.888419 -0.055204
N 11.823496 31.255002 2.384659
N 9.292831 30.783362 3.568730
N 8.036805 31.418327 1.124265
C 9.784765 32.177349 -1.158798
C 10.612656 32.501029 -2.296868
C 11.903375 32.404043 -1.876832
C 11.859093 32.028943 -0.483750
C 12.965737 31.464698 1.641427
C 14.146517 31.236323 2.440231
C 13.713061 30.885870 3.681911
C 12.268752 30.896411 3.634891
C 10.067717 30.486167 4.664747
C 9.246224 30.053411 5.772052
C 7.957075 30.082846 5.336488
C 7.995710 30.538421 3.967046
C 6.900258 31.104497 1.836595
C 5.722470 31.251707 1.015333
C 6.148430 31.668586 -0.207993
C 7.587039 31.767438 -0.130483
C 8.399453 32.134197 -1.192329
H 7.912872 32.388031 -2.131079
C 12.984883 31.836053 0.306093
H 13.955948 31.977044 -0.162626
C 11.453768 30.560663 4.708020
H 11.940677 30.298823 5.644352
C 6.877071 30.697580 3.164102
H 5.907240 30.476797 3.603674
H 12.813946 32.569160 -2.441577
H 10.236332 32.758110 -3.280309
H 15.164312 31.335191 2.080201
H 14.299625 30.629109 4.556760
H 9.626524 29.758225 6.743433
H 7.053076 29.823583 5.875809
H 4.709768 31.058315 1.350561
H 5.561898 31.886355 -1.093106
N 9.832739 33.209042 2.298783
O 9.346337 34.075996 1.606023
"""
geometry_2_00 = """
Fe 9.917990 31.445558 1.778346
N 10.556809 31.866188 -0.055498
N 11.814089 31.227003 2.372666
N 9.297875 30.758246 3.550104
N 8.043584 31.397768 1.120485
C 9.784831 32.164652 -1.160219
C 10.611624 32.501801 -2.293514
C 11.902858 32.406547 -1.875160
C 11.859552 32.017818 -0.486307
C 12.960503 31.454432 1.636717
C 14.140770 31.242960 2.439615
C 13.708543 30.884151 3.678983
C 12.266351 30.874173 3.627468
C 10.070264 30.465070 4.655102
C 9.247247 30.053101 5.766681
C 7.958085 30.091201 5.332866
C 7.998432 30.529979 3.958727
C 6.901428 31.093932 1.833807
C 5.723289 31.255057 1.016540
C 6.151314 31.670649 -0.206350
C 7.589736 31.755538 -0.133074
C 8.400230 32.124963 -1.194447
H 7.913264 32.386655 -2.130914
C 12.983905 31.827747 0.302415
H 13.955696 31.979687 -0.161365
C 11.454251 30.533644 4.698234
H 11.941002 30.276716 5.636156
C 6.877444 30.689985 3.159940
H 5.907605 30.480118 3.604825
H 12.813105 32.581608 -2.437367
H 10.233725 32.768337 -3.273979
H 15.157796 31.357524 2.082132
H 14.295001 30.638320 4.557047
H 9.626721 29.768762 6.741623
H 7.051752 29.847502 5.875478
H 4.709710 31.071712 1.354640
H 5.565103 31.898376 -1.089333
N 9.840508 33.353531 2.373019
O 9.344561 34.158205 1.637232
"""
geometry_5_00 = """
Fe 9.918629 31.289202 1.717339
N 10.542914 31.832173 -0.080685
N 11.795572 31.199413 2.341831
N 9.294593 30.741247 3.513929
N 8.042689 31.359481 1.087282
C 9.775254 32.111817 -1.200449
C 10.600219 32.479101 -2.319680
C 11.891090 32.425876 -1.887580
C 11.847694 32.024341 -0.507342
C 12.945734 31.464689 1.611366
C 14.116395 31.289997 2.423572
C 13.685777 30.915122 3.663719
C 12.252381 30.861042 3.608186
C 10.062170 30.463021 4.634102
C 9.236749 30.104333 5.755782
C 7.945687 30.161198 5.324720
C 7.989641 30.552269 3.941498
C 6.892881 31.087489 1.815829
C 5.722676 31.253502 1.001149
C 6.153153 31.631057 -0.238233
C 7.586010 31.695401 -0.179773
C 8.390724 32.047572 -1.247553
H 7.903308 32.291586 -2.187969
C 12.973334 31.849872 0.283741
H 13.944682 32.031190 -0.169145
C 11.447158 30.518591 4.678739
H 11.934423 30.277429 5.619969
C 6.864795 30.711643 3.146118
H 5.893357 30.532078 3.599511
H 12.800139 32.636412 -2.439296
H 10.224017 32.743662 -3.301293
H 15.131785 31.441247 2.076257
H 14.273933 30.694315 4.546802
H 9.612512 29.848040 6.739754
H 7.036117 29.960530 5.879248
H 4.707408 31.099933 1.347803
H 5.564992 31.851940 -1.121294
N 9.666041 36.091609 3.085945
O 9.598728 37.226756 3.411299
"""
str_geometries = {
"1.75": geometry_1_75,
"2.00": geometry_2_00,
"5.00": geometry_5_00,
}
hivqe_result = {}
{'5.0': '\nFe 9.918629 31.289202 1.717339\nN 10.542914 31.832173 -0.080685\nN 11.795572 31.199413 2.341831\nN 9.294593 30.741247 3.513929\nN 8.042689 31.359481 1.087282\nC 9.775254 32.111817 -1.200449\nC 10.600219 32.479101 -2.319680\nC 11.891090 32.425876 -1.887580\nC 11.847694 32.024341 -0.507342\nC 12.945734 31.464689 1.611366\nC 14.116395 31.289997 2.423572\nC 13.685777 30.915122 3.663719\nC 12.252381 30.861042 3.608186\nC 10.062170 30.463021 4.634102\nC 9.236749 30.104333 5.755782\nC 7.945687 30.161198 5.324720\nC 7.989641 30.552269 3.941498\nC 6.892881 31.087489 1.815829\nC 5.722676 31.253502 1.001149\nC 6.153153 31.631057 -0.238233\nC 7.586010 31.695401 -0.179773\nC 8.390724 32.047572 -1.247553\nH 7.903308 32.291586 -2.187969\nC 12.973334 31.849872 0.283741\nH 13.944682 32.031190 -0.169145\nC 11.447158 30.518591 4.678739\nH 11.934423 30.277429 5.619969\nC 6.864795 30.711643 3.146118\nH 5.893357 30.532078 3.599511\nH 12.800139 32.636412 -2.439296\nH 10.224017 32.743662 -3.301293\nH 15.131785 31.441247 2.076257\nH 14.273933 30.694315 4.546802\nH 9.612512 29.848040 6.739754\nH 7.036117 29.960530 5.879248\nH 4.707408 31.099933 1.347803\nH 5.564992 31.851940 -1.121294\nN 9.666041 36.091609 3.085945\nO 9.598728 37.226756 3.411299\n'}
geometry_1_75 = """
Fe 9.910596 31.534095 1.798088
N 10.557481 31.888419 -0.055204
N 11.823496 31.255002 2.384659
N 9.292831 30.783362 3.568730
N 8.036805 31.418327 1.124265
C 9.784765 32.177349 -1.158798
C 10.612656 32.501029 -2.296868
C 11.903375 32.404043 -1.876832
C 11.859093 32.028943 -0.483750
C 12.965737 31.464698 1.641427
C 14.146517 31.236323 2.440231
C 13.713061 30.885870 3.681911
C 12.268752 30.896411 3.634891
C 10.067717 30.486167 4.664747
C 9.246224 30.053411 5.772052
C 7.957075 30.082846 5.336488
C 7.995710 30.538421 3.967046
C 6.900258 31.104497 1.836595
C 5.722470 31.251707 1.015333
C 6.148430 31.668586 -0.207993
C 7.587039 31.767438 -0.130483
C 8.399453 32.134197 -1.192329
H 7.912872 32.388031 -2.131079
C 12.984883 31.836053 0.306093
H 13.955948 31.977044 -0.162626
C 11.453768 30.560663 4.708020
H 11.940677 30.298823 5.644352
C 6.877071 30.697580 3.164102
H 5.907240 30.476797 3.603674
H 12.813946 32.569160 -2.441577
H 10.236332 32.758110 -3.280309
H 15.164312 31.335191 2.080201
H 14.299625 30.629109 4.556760
H 9.626524 29.758225 6.743433
H 7.053076 29.823583 5.875809
H 4.709768 31.058315 1.350561
H 5.561898 31.886355 -1.093106
N 9.832739 33.209042 2.298783
O 9.346337 34.075996 1.606023
"""
geometry_2_00 = """
Fe 9.917990 31.445558 1.778346
N 10.556809 31.866188 -0.055498
N 11.814089 31.227003 2.372666
N 9.297875 30.758246 3.550104
N 8.043584 31.397768 1.120485
C 9.784831 32.164652 -1.160219
C 10.611624 32.501801 -2.293514
C 11.902858 32.406547 -1.875160
C 11.859552 32.017818 -0.486307
C 12.960503 31.454432 1.636717
C 14.140770 31.242960 2.439615
C 13.708543 30.884151 3.678983
C 12.266351 30.874173 3.627468
C 10.070264 30.465070 4.655102
C 9.247247 30.053101 5.766681
C 7.958085 30.091201 5.332866
C 7.998432 30.529979 3.958727
C 6.901428 31.093932 1.833807
C 5.723289 31.255057 1.016540
C 6.151314 31.670649 -0.206350
C 7.589736 31.755538 -0.133074
C 8.400230 32.124963 -1.194447
H 7.913264 32.386655 -2.130914
C 12.983905 31.827747 0.302415
H 13.955696 31.979687 -0.161365
C 11.454251 30.533644 4.698234
H 11.941002 30.276716 5.636156
C 6.877444 30.689985 3.159940
H 5.907605 30.480118 3.604825
H 12.813105 32.581608 -2.437367
H 10.233725 32.768337 -3.273979
H 15.157796 31.357524 2.082132
H 14.295001 30.638320 4.557047
H 9.626721 29.768762 6.741623
H 7.051752 29.847502 5.875478
H 4.709710 31.071712 1.354640
H 5.565103 31.898376 -1.089333
N 9.840508 33.353531 2.373019
O 9.344561 34.158205 1.637232
"""
geometry_5_00 = """
Fe 9.918629 31.289202 1.717339
N 10.542914 31.832173 -0.080685
N 11.795572 31.199413 2.341831
N 9.294593 30.741247 3.513929
N 8.042689 31.359481 1.087282
C 9.775254 32.111817 -1.200449
C 10.600219 32.479101 -2.319680
C 11.891090 32.425876 -1.887580
C 11.847694 32.024341 -0.507342
C 12.945734 31.464689 1.611366
C 14.116395 31.289997 2.423572
C 13.685777 30.915122 3.663719
C 12.252381 30.861042 3.608186
C 10.062170 30.463021 4.634102
C 9.236749 30.104333 5.755782
C 7.945687 30.161198 5.324720
C 7.989641 30.552269 3.941498
C 6.892881 31.087489 1.815829
C 5.722676 31.253502 1.001149
C 6.153153 31.631057 -0.238233
C 7.586010 31.695401 -0.179773
C 8.390724 32.047572 -1.247553
H 7.903308 32.291586 -2.187969
C 12.973334 31.849872 0.283741
H 13.944682 32.031190 -0.169145
C 11.447158 30.518591 4.678739
H 11.934423 30.277429 5.619969
C 6.864795 30.711643 3.146118
H 5.893357 30.532078 3.599511
H 12.800139 32.636412 -2.439296
H 10.224017 32.743662 -3.301293
H 15.131785 31.441247 2.076257
H 14.273933 30.694315 4.546802
H 9.612512 29.848040 6.739754
H 7.036117 29.960530 5.879248
H 4.707408 31.099933 1.347803
H 5.564992 31.851940 -1.121294
N 9.666041 36.091609 3.085945
O 9.598728 37.226756 3.411299
"""
str_geometries = {
"1.75": geometry_1_75,
"2.00": geometry_2_00,
"5.00": geometry_5_00,
}
hivqe_result = {}
{'5.0': '\nFe 9.918629 31.289202 1.717339\nN 10.542914 31.832173 -0.080685\nN 11.795572 31.199413 2.341831\nN 9.294593 30.741247 3.513929\nN 8.042689 31.359481 1.087282\nC 9.775254 32.111817 -1.200449\nC 10.600219 32.479101 -2.319680\nC 11.891090 32.425876 -1.887580\nC 11.847694 32.024341 -0.507342\nC 12.945734 31.464689 1.611366\nC 14.116395 31.289997 2.423572\nC 13.685777 30.915122 3.663719\nC 12.252381 30.861042 3.608186\nC 10.062170 30.463021 4.634102\nC 9.236749 30.104333 5.755782\nC 7.945687 30.161198 5.324720\nC 7.989641 30.552269 3.941498\nC 6.892881 31.087489 1.815829\nC 5.722676 31.253502 1.001149\nC 6.153153 31.631057 -0.238233\nC 7.586010 31.695401 -0.179773\nC 8.390724 32.047572 -1.247553\nH 7.903308 32.291586 -2.187969\nC 12.973334 31.849872 0.283741\nH 13.944682 32.031190 -0.169145\nC 11.447158 30.518591 4.678739\nH 11.934423 30.277429 5.619969\nC 6.864795 30.711643 3.146118\nH 5.893357 30.532078 3.599511\nH 12.800139 32.636412 -2.439296\nH 10.224017 32.743662 -3.301293\nH 15.131785 31.441247 2.076257\nH 14.273933 30.694315 4.546802\nH 9.612512 29.848040 6.739754\nH 7.036117 29.960530 5.879248\nH 4.707408 31.099933 1.347803\nH 5.564992 31.851940 -1.121294\nN 9.666041 36.091609 3.085945\nO 9.598728 37.226756 3.411299\n'}
Passo 2 e 3: Otimizar o problema para execução em hardware quântico e executar usando a função HiVQE Chemistry
Com base na configuração do HiVQE e das geometrias, obtenha os resultados sequencialmente.
Submeter o cálculo d(Fe-N) = 1,75 .
hivqe_run_1_75 = hivqe.run(
geometry=str_geometries["1.75"],
backend_name="",
max_states=400000000,
max_expansion_states=100,
molecule_options=molecule_options,
hivqe_options=hivqe_options,
)
info_jobid_1_75 = hivqe_run_1_75.job_id
Acompanhe o job e recupere o resultado para o cálculo d(Fe-N) = 1,75 .
submitted_job_1_75 = catalog.get_job_by_id(info_jobid_1_75)
stat = submitted_job_1_75.status()
print(submitted_job_1_75.job_id, stat)
if stat == "DONE":
hivqe_run_1_75_energy = submitted_job_1_75.result()["energy"]
print(f"Completed HiVQE calculation, Energy {hivqe_run_1_75_energy}")
hivqe_result["1.75"] = hivqe_run_1_75_energy
Submeter o cálculo d(Fe-N) = 2,00 .
hivqe_run_2_00 = hivqe.run(
geometry=str_geometries["2.00"],
backend_name="",
max_states=400000000,
max_expansion_states=100,
molecule_options=molecule_options,
hivqe_options=hivqe_options,
)
info_jobid_2_00 = hivqe_run_2_00.job_id
Acompanhe o job e recupere o resultado para o cálculo d(Fe-N) = 2,00 .
submitted_job_2_00 = catalog.get_job_by_id(info_jobid_2_00)
stat = submitted_job_2_00.status()
print(submitted_job_2_00.job_id, stat)
if stat == "DONE":
hivqe_run_2_00_energy = submitted_job_2_00.result()["energy"]
print(f"Completed HiVQE calculation, Energy {hivqe_run_2_00_energy}")
hivqe_result["2.00"] = hivqe_run_2_00_energy
Submeter o cálculo d(Fe-N) = 5,00 .
hivqe_run_5_00 = hivqe.run(
geometry=str_geometries["5.00"],
backend_name="",
max_states=400000000,
max_expansion_states=100,
molecule_options=molecule_options,
hivqe_options=hivqe_options,
)
info_jobid_5_00 = hivqe_run_5_00.job_id
Acompanhe o job e recupere o resultado para o cálculo d(Fe-N) = 5,00 .
submitted_job_5_00 = catalog.get_job_by_id(info_jobid_5_00)
stat = submitted_job_5_00.status()
print(submitted_job_5_00.job_id, stat)
if stat == "DONE":
hivqe_run_5_00_energy = submitted_job_5_00.result()["energy"]
print(f"Completed HiVQE calculation, Energy {hivqe_run_5_00_energy}")
hivqe_result["5.00"] = hivqe_run_5_00_energy
hivqe_result = {
"1.75": -2373.681781,
"2.00": -2373.694128,
"5.00": -2373.637807,
}
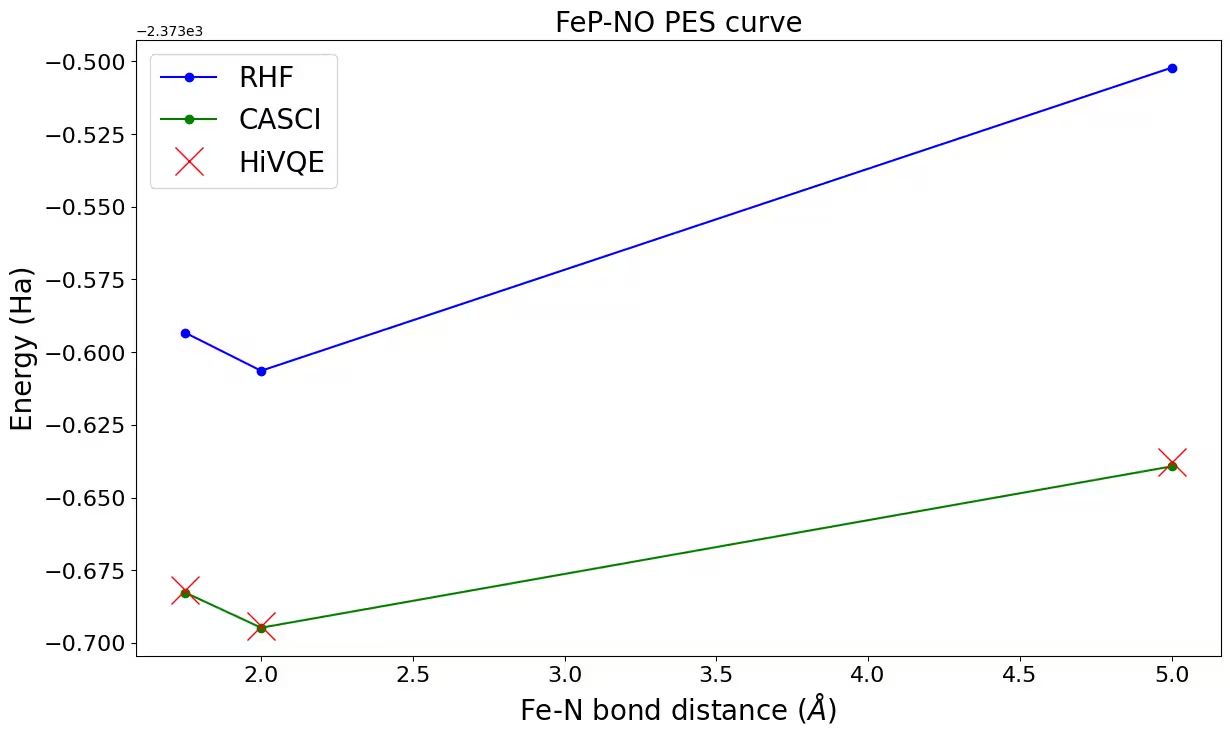
Passo 4: Pós-processar e comparar com métodos clássicos
Os resultados de cálculo de referência clássico (CASCI-DMRG, maxM=800) são fornecidos para (22o,22e) para validar os resultados do HiVQE.
rhf_result = {
"1.75": -2373.59331683504,
"2.00": -2373.60640773065,
"5.00": -2373.50214278007,
}
casci_result = {"1.75": -2373.6827, "2.00": -2373.6948, "5.00": -2373.6393}
fig, ax = plt.subplots(1, 1)
hf_energy = [v for key, v in rhf_result.items()]
casci_energy = [v for key, v in casci_result.items()]
hivqe_energy = [v for key, v in hivqe_result.items()]
distance_ref = [float(key) for key, v in rhf_result.items()]
distance = [float(key) for key, v in hivqe_result.items()]
ax.plot(distance_ref, hf_energy, "-o", label="RHF", c="blue")
ax.plot(distance_ref, casci_energy, "-o", label="CASCI", c="green")
ax.plot(distance, hivqe_energy, "x", label="HiVQE", c="red", markersize=20)
ax.legend(fontsize=20)
ax.tick_params("both", labelsize=16)
ax.set_xlabel("Fe-N bond distance ($\AA$)", size=20)
ax.set_ylabel("Energy (Ha)", size=20)
ax.set_title("FeP-NO PES curve", size=20)
fig.set_size_inches(14, 8)
Pesquisa do tutorial
Por favor, responda a esta breve pesquisa para fornecer feedback sobre este tutorial. Seus insights nos ajudarão a melhorar nossas ofertas de conteúdo e experiência do usuário.
Note: This survey is provided by IBM Quantum and relates to the original English content. To give feedback on doQumentation's website, translations, or code execution, please open a GitHub issue.